WISP
An optimized algorithm and tool for dynamical network analysis
Introduction
Allostery can occur by way of subtle cooperation among protein residues (e.g., amino acids) even in the absence of large conformational shifts. Dynamical network analysis has been used to model this cooperation, helping to computationally explain how binding to an allosteric site can impact the behavior of a primary site often many angstroms away. Traditionally, computational efforts have focused on the most optimal path of correlated motions leading from the allosteric to the primary active site. We present a program called Weighted Implementation of Suboptimal Paths (WISP) capable of rapidly identifying additional suboptimal pathways that may also play important roles in the transmission of allosteric signals. Aside from providing signal redundancy, suboptimal paths traverse residues that, if disrupted through pharmacological or mutational means, could modulate the allosteric regulation of important drug targets.
Download
WISP and it's easy-to-use VMD plugin can be downloaded free of charge from sourceforge.net. The program is released under the Academic Free License 3.0 license.
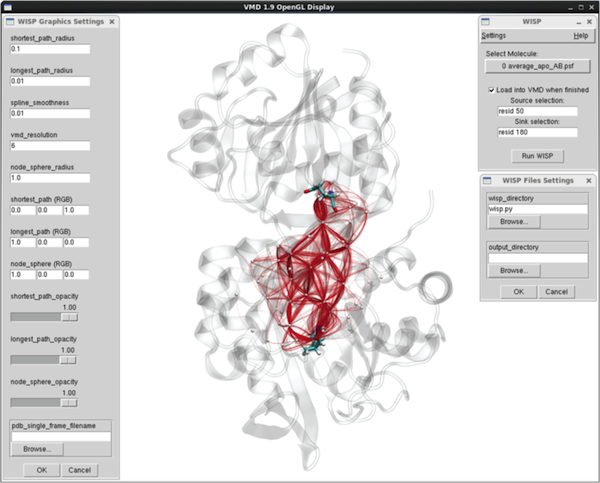
The WISP VMD-plugin GUI.
Example Output

The WISP VMD-plugin GUI.
Usage
FILE-SYSTEM PARAMETERS
-
pdb_trajectory_filename: - The filename of the multi-frame PDB to analyze. Individual frames should be separated by "END" or "ENDMDL" lines.
-
output_directory: -
A new directory where the WISP output should be
written. If this parameter is not specified, a default output
directory is created whose name includes the current date for
future reference. The default value is
wisp_output__Sep_17_2013__10_44_PM.
COVARIANCE-MATRIX PARAMETERS
-
node_definition: - WISP calculates the covariance matrix by defining nodes associated with each protein residue. If node_definition is set to "CA," the alpha carbon will be used. If set to "RESIDUE_COM,", "SIDECHAIN_COM,", or "BACKBONE_COM," the whole- residue, side-chain, or backbone center of mass will be used, respectively. The default value is RESIDUE_COM.
-
contact_map_distance_limit: - If you use WISP's default contact-map generator, node pairs with average inter-node distances greater than this value will not be considered in calculating the covariance matrix. The default value is 999999.999.
-
load_wisp_saved_matrix: - If the covariance matrix (appropriately modifed by a contact map) has been previously saved to a file, set this parameter to "TRUE" to load the matrix instead of generating it from scratch. WISP automatically saves a copy of this matrix to the file "functionalized_matrix_with_contact_map_applied.pickle" in the output directory every time it is run. The default value is FALSE.
-
wisp_saved_matrix_filename: - If load_wisp_saved_matrix is set to "TRUE," this parameter specifies the file to load. If it is set to "FALSE," this parameter specifies the file to which the matrix should be saved.
PATH-SEARCHING PARAMETERS
-
desired_number_of_paths: - One of the advantages of WISP is that it can calculate not only the optimal path between residues, but multiple good paths. This parameter specifies the desired number of paths. The default value is 1.
-
source_residues: - This parameter specifies the source residues for path generation. A list of residues should be constructed of the form "CHAIN_RESNAME_RESID," separated by spaces. For example: "X_SER_1 X_LEU_4." For unix to treat a space-containing command-line parameter as a single parameter, it must be enclosed in quotes.
-
sink_residues: - This parameter specifies the sink residues for path generation. The format is the same as for the source_residues parameter.
MULTI-PROCESSOR PARAMETERS
-
number_processors: - On unix-like machines, WISP can use multiple processors to significantly increase speed. This parameter specifies the number of processors to use. The default value is 1.
-
num_frames_to_load_before_processing: - When WISP is run with multiple processors, the frames from the PDB are loaded in chunks before being distributed to the many processors. This parameter specifies the number of frames to load before distribution. The default value is 96.
VISUALIZATION PARAMETERS
-
shortest_path_radius: - WISP outputs a VMD state file to facilitate visualization. The shortest path is represented by a strand with the largest radius. Longer paths have progressively smaller radii. This parameter specifies the radius of the shortest path, in Angstroms. The default value is 0.1.
-
longest_path_radius: - This parameter specifies the radius of the longest path visualized, in Angstroms. The default value is 0.01.
-
spline_smoothness: - The paths are represented by splines connecting the nodes. This parameter indicates the smoothness of the splines. Smaller values produce smoother splies, but take longer to render. The default value is 0.01.
-
vmd_resolution: - When visualizing in VMD, a number of cylinders and spheres are drawn. This parameter specifies the resolution to use. The default value is 6.
-
node_sphere_radius: - When visualizing in VMD, spheres are placed at the locations of the nodes. This parameter specifies the radius of these spheres. The default value is 1.0.
-
shortest_path_r: - The color of the shortest path is given by an RGB color code. This parameter specifies the R value, ranging from 0.0 to 1.0. The default value is 0.0.
-
shortest_path_g: - The color of the shortest path is given by an RGB color code. This parameter specifies the G value, ranging from 0.0 to 1.0. The default value is 0.0.
-
shortest_path_b: - The color of the shortest path is given by an RGB color code. This parameter specifies the B value, ranging from 0.0 to 1.0. The default value is 1.0.
-
longest_path_r: - The color of the longest path is given by an RGB color code. This parameter specifies the R value, ranging from 0.0 to 1.0. The default value is 1.0.
-
longest_path_g: - The color of the longest path is given by an RGB color code. This parameter specifies the G value, ranging from 0.0 to 1.0. The default value is 0.0.
-
longest_path_b: - The color of the longest path is given by an RGB color code. This parameter specifies the B value, ranging from 0.0 to 1.0. The default value is 0.0.
-
node_sphere_r: - The color of the node spheres is given by an RGB color code. This parameter specifies the R value, ranging from 0.0 to 1.0. The default value is 1.0.
-
node_sphere_g: - The color of the node spheres is given by an RGB color code. This parameter specifies the G value, ranging from 0.0 to 1.0. The default value is 1.0.
-
node_sphere_b: - The color of the node spheres is given by an RGB color code. This parameter specifies the B value, ranging from 0.0 to 1.0. The default value is 1.0.
-
shortest_path_opacity: - The opacity of the shortest path, ranging from 0.0 (transparent) to 1.0 (fully opaque). Note that if --shortest_path_opacity, --longest_path_opacity, and --node_sphere_opacity are not all identical, the output TCL file will contain many materials, which may be less-than-desirable for some users. The default value is 1.0.
-
longest_path_opacity: - The opacity of the longest path, ranging from 0.0 (transparent) to 1.0 (fully opaque). The default value is 1.0.
-
node_sphere_opacity: - The opacity of the node spheres, ranging from 0.0 (transparent) to 1.0 (fully opaque). The default value is 1.0.
-
pdb_single_frame_filename: - By default, WISP uses the trajectory- average structure for positioning the nodes, visualizing the paths and protein, etc. However, if desired, a separate PDB structure with the same residue order and number can be specified for this purpose using the "pdb_single_frame_filename" parameter.
ADVANCED FEATURES
-
seconds_to_wait_before_parallelizing_path_finding: - WISP identifies paths from the source to the sink by recursively visiting node neighbors. The program begins the recursion algorithm on a single processor before distributing the search efforts to multiple processors. This parameter specifies how long WISP should search for source-sink paths using a single processor before distributing the search effort over multiple processors. By waiting longer before distribution, the search efforts are ultimately distributed more evenly over the multiple processors, potentially increasing speed in the long run. On the other hand, specifiying a lower value for this parameter means the program will spend more time running on multiple processors, also potentially increasing speed. A balance must be struck. The default value is 5.0.
-
user_specified_functionalized_matrix_filename: - A text file containing a user-specified functionalized correlation matrix. If not given, WISP's default functionalized correlation matrix, as described in the WISP publication, will be automatically calculated. For convenience, WISP automatically saves a human-readable copy of the matrix used to the file "functionalized_correlation_matrix.txt" in the output directory every time it is run.
-
user_specified_contact_map_filename: - A text file containing a user- specified contact map. If given, each element of the functionalized matrix will be multiplied by the corresponding value specified in the file. If not given, WISP's default contact map, based on the distances between average node locations, will be automatically applied. For convenience, WISP automatically saves a human-readable copy of the contact-map matrix to the file "contact_map_matrix.txt" in the output directory every time it is run.
Notes:
- To visualize in VMD, first load the output TCL file, then load the PDB file.
- WISP ignores PDB segnames. Every residue in your PDB trajectory must be uniquely identifiable by the combination of its chain, resname, and resid.
Example:
python wisp.py
-pdb_trajectory_filename multi_frame_pdb.pdb
-node_definition CA
-contact_map_distance_limit 50.0
-load_wisp_saved_matrix false
-wisp_saved_matrix_filename matrix.file
-desired_number_of_paths 30
-source_residues "X_SER_1 X_LEU_4"
-sink_residues X_ARG_37
-number_processors 24
-num_frames_to_load_before_processing 96
-seconds_to_wait_before_parallelizing_path_finding 10.0
-shortest_path_radius 0.2
-longest_path_radius 0.05
-spline_smoothness 0.05
-vmd_resolution 6
-node_sphere_radius 1.0
FTProd is freely available under the GNU public license. If you have any suggestions, issues or questions, please don't hesitate to contact me, Lane Votapka, at lvotapka100@gmail.com.