Single Frame Electrostatics
Loading the molecule:
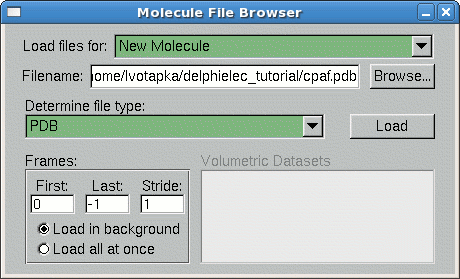
- On the VMD main window, go to File → New Molecule
- The "Molecule File Browser" window will appear. Under the "Load Files for:" list box, select "New Molecule"
- Click "Browse…"
- Navigate to "…/cpaf.pdb" Click "OK"
- In the Molecule File Browser, Click "Load"
Start the Delphi run
- In the VMD Main window, select Extensions → Analysis → DelEnsembleElec. The DelEnsembleElec plugin window will appear.
- Under the option box "Select molecule for Delphi run:" select "0:cpaf.pdb"
- In the "Selection" textbox, change the value to "protein"
- In the "Write to Output" textbox, change the value to "struct_cpaf.cube"
- Make sure the "Load into VMD when finished" option is selected.
- Go to the menubar "Edit" → "Input File". The Edit Delphi Input File window will appear.
- Click the button "Load Template Param. File"
- Navigate to the file "…/param.in". Click "Open"
- The "Size File" and "Charge File" fields must point to the proper .siz and .crg parameter files that is included in this tutorial or may be downloaded at Clemson's Website, and should be under the "Parameter Files" tab.
- Click "OK" to close the Edit Delphi Input File window
-
Click the "Run Delphi" button.
Note: This may take several minutes to complete.

Examine the results
Once finished, examine the results of the run:
- Try changing the "Isovalue" slider in the Graphical Representations window under the "Draw style" tab
- Try changing the Drawing Method listbox to another value: such as "Volume Slice" or "Field Lines"
- Try creating a positive and negative representation of the charge maps by clicking the "Create Rep" button in the Graphical Representations window and changing the isovalue slider to a positive number, and the isovalue of the first representation to a negative number.

Next
Plugin Developer
Lane Votapka
Contributors
Rommie Amaro, Luke Czapla, Maxim
Zhenirovskyy