Strain Comparison
To begin, we will use FTProd to compare three different strains of neuraminidase. Neuraminidase is an enzyme found on the outer envelope of the influenza virus. It normally exists as a tetramer, but we will only consider the monomer form in this tutorial. We have already run the PDB structures 1MWE, 2HU0, and 3NSS through FTMAP for you, but if you want to, you can run them through FTMAP yourself by visiting the FTMAP server", creating an account, and entering these PDB codes in the 'Submit' page.
Preparing tutorial files
- Download the tutorial files
-
Unzip by clicking on it or with this command:
tar -xfz ftprod_tutorial.tgz
Loading the molecules
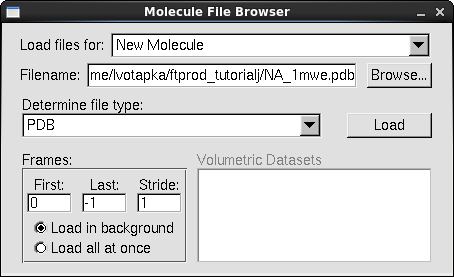
- In a fresh session of VMD, in the VMD main window, go to File → New Molecule. The "Molecule File Browser" window will appear.
- Under the "Load Files for:" list box, select "New Molecule"
- Click "Browse…"
- Navigate to the tutorial directory and select the file"NA_1mwe.pdb" Click "OK"
- In the Molecule File Browser, Click "Load"
- Repeat the above instructions for the files: "NA_2hu0.pdb" & "NA_3nss.pdb", making sure to select "New Molecule" option under "Load files for"
Align the Molecules
Note that the molecules aren't aligned. Equivalent FTMAP structures must be superimposed for FTProd to be able to identify respective consensus sites
- In the VMD main window, go to Extensions → Analysis → MultiSeq. The MultiSeq window should appear
- In the MultiSeq window table where the protein structures' names are listed, select the three checkboxes next to the names of our proteins: NA_1mwe_A, NA_2hu0_A, and NA_3nss_A
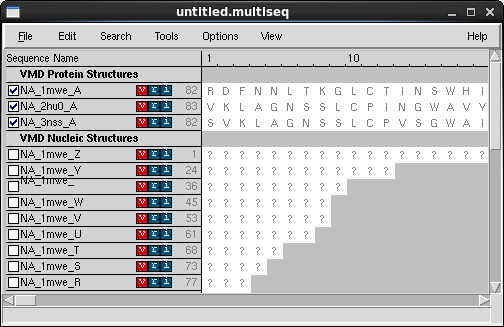
- In the MultiSeq menu, select Tools → Stamp Structural Alignment. The Stamp Alignment Options window should appear.
- In the "Align the Following" frame, select the "Marked Structures" option.
- In the "Similarity (scanscore)" area, move the slider down from 6 to 5
- Leave the other options at their defaults and click OK
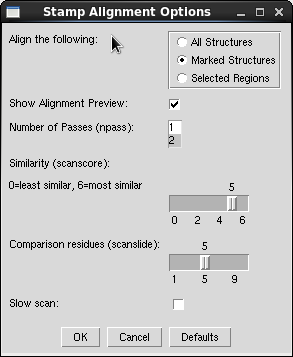

- Within a moment, the neuraminidase structures should align in the VMD view window.
Run FTProd
-
In the VMD main window, go to Extensions → Analysis → FTProd. The FTProd main window should
appear.
All loaded molecules appear in the "Selected Molecule(s)" listbox. The three loaded structures should be listed there, as well as any others you may have loaded. -
In the "Selected Molecule(s)" listbox, select the three neuraminidase structures. Or alternatively,
click the "Select All" button below the list to select all loaded structures.
The "CS distance cutoff" slider allows you to specify the distance beyond which consensus sites are no longer clustered. A larger CS distance cutoff will result in larger clusters and vice versa. -
Set the CS distance cutoff to 7.0.
You can move the slider with your mouse or enter the number in the adjacent textbox manually - Leave the other options at their defaults and click "Run"
Examine the Results
- The "FTProd Table" window will appear
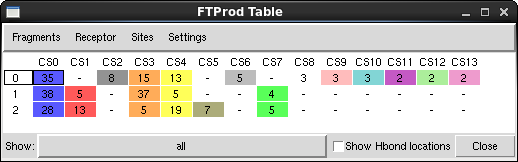
NOTE: if you ever accidentally close the Table window, you can reopen it again my going to Views → Table in the FTProd main window.
- Click on some of the colored rectangles in the table window. This allows you to hide and show the Representations you wish to view
- The three structures are located along the left side of the table. The consensus sites identified by FTProd are located along the top. Notice that some Consensus sites are found in many structures, and some are only found in one or two
-
Click on the first Consensus site (CS0) for the NA_1mwe.pdb structure (the first blue rectangle). This is neuraminidase's sialic acid binding site, and is also the site that binds to the neuraminidase inhibitor Tamiflu.
You can click and drag, or Shift-click or Ctrl-click to select multiple consensus sites.
- Click the next blue Consensus site right below the first. The structure will change to the sialic acid binding site of NA_2hu0. Notice that FTProd has classified these two sites as equivalent because of their spatial proximity.
- Try clicking on some of the other sites. If you click on a red rectangle in the second column, a different consensus site will be highlighted. Notice that it only exists in the structures NA_2hu0 and NA_3nss.
-
If you click the grey rectangle in the third column, it will highlight a consensus site that exists only in structure NA_1mwe
FTProd also allows you to easily modify the appearance of the consensus site, its associated fragments, and the protein (receptor).
- In the FTProd Table window menu, go to Sites → Coloring method → ResType. All consensus site residues are now colored according to VMD's ResType coloring scheme.
- Next, in the FTProd Table window menu, go to Fragment → Drawing method → VDW. All probe molecules will now be drawn as Van der Waals spheres.
- Then, in the FTProd Table window menu, go to Sites → Drawing method → Licorice. All consensus site residues are now drawn as licorice
- In the FTProd Table window, beside the "Show" label, click the option menu that currently says "all" and instead select "H-bond donor". Now only probe molecules classified as hydrogen bond donors will be displayed.
Other features
-
Fragments can be colored by hydrophobicity. This is done by going to Fragments → Coloring method → Hydrophobic. The atoms are then colored on a scale from white to green. The greener the atom, the more hydrophobic it is.
How does FTProd get this number? It does it by placing a 3-D gaussian function over every heavy hydrophobic probe atom. These gaussian functions are summed, and then the value is calculated for the locations of EVERY probe atom. - FTProd can also display the locations where multiple probe hydrogen bond donors/acceptors appear. Toggle the checkbox labelled "Show Hbond locations" in the FTProd Table window. Many blue and red icosahedrons will appear. A red icosahedron indicates a location where two or more H-bond acceptors appear. A blue icosahedron indicates a H-bond donor. A red and blue icosahedron indicates where both exist in relatively equal quantities. The H-bond icosahedrons also vary by size. A large icosahedron indicates a location where many H-bond donors/acceptors appear and vice versa.
Next
Plugin Developer
Lane Votapka
Contributors
Rommie Amaro, Ozlem Demir, Rob Swift,
Robert Malmstrom
Testers
Rachel Li, Francesca Bardinelli, Pek Ieong, Emilia Pecora de
Barros, Sophia Hirakis, Saira Ikram, Eric Chen